New York City’s emergency rooms are straining under what public health officials are calling the worst flu surge in a decade. During the week ending December 20, nearly 10,000 patients flooded city emergency departments complaining of flu-like symptoms—more than during any comparable week in the past ten years. At Long Island Jewish Medical Center in Manhasset, daily emergency room visits have jumped from 250 to nearly 290, with the dominant H3N2 strain showing mutations that help it evade immune defenses. Schools have shuttered, including Brooklyn’s Poly Prep Country Day School, where roughly a third of students fell ill within days. And public health experts warn the worst is yet to come.
Against this backdrop, a new study published in the Proceedings of the National Academy of Sciences offers a remarkable window into the microscopic battle occurring in our bodies millions of times over this holiday season. Researchers from Switzerland and Japan have, for the first time, captured live, high-resolution video of influenza viruses entering living human cells—revealing a process far more dynamic and surprising than scientists previously understood.
“The infection of our body cells is like a dance between virus and cell,” says Yohei Yamauchi, Professor of Molecular Medicine, ETH Zurich.
That observation, from lead researcher Yohei Yamauchi, upends the conventional view of viral infection as a one-sided assault. The study demonstrates that human cells are not passive victims simply overwhelmed by invading pathogens. Instead, they actively participate in their own infection—a finding with profound implications for developing new antiviral therapies.

A Virus-Eye View of Infection
The breakthrough centers on a novel imaging system the researchers call ViViD-AFM, which combines atomic force microscopy with confocal fluorescence microscopy. Previous attempts to study viral entry faced a fundamental limitation: electron microscopy techniques that offered sufficient resolution required destroying the cells being studied, providing only static snapshots of a dynamic process. Fluorescence microscopy, while allowing live observation, lacked the spatial resolution to capture nanoscale details.
The team’s solution was elegant: they redesigned the cantilever—the tiny probe that scans surfaces in atomic force microscopy—making it ultra-narrow with less than half the spring constant of conventional designs. This seemingly simple modification reduced the mechanical force exerted during scanning from 25 piconewtons to just 10 piconewtons, matching the lower threshold of the weak bonds holding influenza viruses to cell surfaces. The result was an imaging system gentle enough to observe virus-cell interactions without disrupting them.
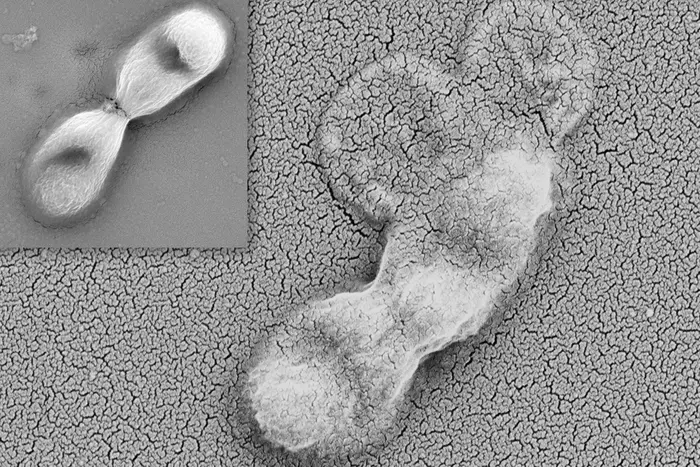
“Thanks to this method, it is now possible to follow the detailed dynamics of the virus’s entry into the cell,” the researchers noted. Using fluorescently labeled influenza A viruses, they tracked individual viral particles as they landed on living kidney cells—the same cell type commonly used in influenza research due to its high susceptibility to infection.

Surfing Before Sinking
What the researchers observed challenges textbook descriptions of viral entry. Influenza viruses don’t simply land and bore into cells. Instead, they “surf” across the cell surface, repeatedly attaching and detaching from sugar molecules called sialic acids that stud the outer membrane. This lateral diffusion allows the virus to scan the cellular landscape, searching for optimal entry points where receptor molecules cluster together.
The dynamics are governed by two viral surface proteins working in opposition. Hemagglutinin binds to sialic acids, tethering the virus to the cell. Neuraminidase cleaves these same sugars, releasing the virus to continue its search. This molecular push-and-pull creates the observed “surfing” behavior. When the researchers blocked neuraminidase activity with oseltamivir—the active compound in Tamiflu—viral mobility decreased significantly, demonstrating how the drug may slow infection even before the virus enters cells.
The virus eventually locates receptor-rich domains and triggers a cellular response. The cell actively recruits clathrin proteins to the viral attachment site, forming a pocket that begins engulfing the particle. As this pit deepens, the cell generates actin-rich membrane bulges that rise approximately 100 nanometers high, actively wrapping around the virus and pulling it inside. The whole process culminates in complete membrane envelopment at peak clathrin accumulation.
“The cell actively recruits the functionally important clathrin proteins to the point where the virus is located. The cell surface also actively captures the virus by bulging up at the point in question.”

From Bench to Bedside
The findings arrive at a critical moment. This season’s dominant influenza strain, an H3N2 variant known as subclade K, has acquired mutations that reduce vaccine effectiveness to an estimated 32-39 percent in adults—well below the typical 40-60 percent range. New York State has recorded its highest number of flu cases in a single week since tracking began in 2004, with over 71,000 cases reported. The virus’s genetic drift from vaccine predictions underscores the urgent need for novel therapeutic approaches.
Understanding precisely how influenza enters cells opens multiple avenues for intervention. The researchers demonstrated that their system can distinguish between viral subtypes—H1N1 and H3N2 showed statistically different diffusion coefficients—potentially allowing rapid assessment of how mutations affect viral behavior. This capability could prove invaluable for evaluating emerging variants.
“The new technique therefore provides key insights when it comes to the development of antiviral drugs,” the researchers wrote. “For example, it is suitable for testing the efficacy of potential drugs in a cell culture in real time.” They emphasize the technique could also be applied to study other viruses or even vaccine delivery mechanisms using lipid nanoparticles.

Beyond Influenza
The implications extend beyond the flu. Influenza’s interactions with cell membranes are among the weakest of any virus, with binding forces measuring just 10-25 piconewtons. The system’s success with such delicate interactions suggests it could visualize the entry mechanisms of virtually any virus, including those with stronger receptor affinities.
As New Yorkers crowd emergency rooms this holiday season, many clutching tissues and battling fevers, the microscopic drama playing out in their cells follows the ancient choreography the Japanese and Swiss researchers have now illuminated. Each of those nearly 10,000 weekly emergency visits represents countless individual cellular battles—viruses surfing, cells responding, membranes bulging and engulfing.
“We haven’t hit the peak yet,” warned Caitlin Rivers, an infectious-disease epidemiologist at Johns Hopkins Bloomberg School of Public Health. But in laboratories across the world, scientists now have an unprecedented view of the enemy at the gates—and perhaps, in time, new weapons to turn back the invasion.
Endnotes
1. Yoshida A, Uekusa Y, Suzuki T, Bauer M, Sakai N, Yamauchi Y. Enhanced visualization of influenza A virus entry into living cells using virus-view atomic force microscopy. PNAS. 2025;122(38):e2500660122. doi:10.1073/pnas.2500660122
2. ETH Zurich press release: “How influenza viruses enter our cells.” EurekAlert!, December 4, 2025.
3. New York Times. “‘Super Flu’ Sends NYC Flu Cases to Highest Level in a Decade.” December 26, 2025.
4. Becker’s Hospital Review. “Flu rates jump in New York City, Boston and Texas: What to know.” December 26, 2025.
5. New York State Department of Health press release: “New York State Department of Health Declares Flu Is Prevalent Across the State.” December 18, 2025.
6. NYC Health Department press release: “Don’t Miss Out This Holiday Season, Stay Protected Against COVID-19, Flu, and RSV.” December 2025.
7. Rust MJ, Lakadamyali M, Zhang F, Zhuang X. Assembly of endocytic machinery around individual influenza viruses during viral entry. Nature Structural & Molecular Biology. 2004;11(6):567-573.
8. Sieben C, Sezgin E, Eggeling C, Manley S. Influenza A viruses use multivalent sialic acid clusters for cell binding and receptor activation. PLoS Pathogens. 2020;16(7):e1008656.
9. de Vries E, et al. Dissection of the influenza A virus endocytic routes reveals macropinocytosis as an alternative entry pathway. PLoS Pathogens. 2011;7(3):e1001329.



Leave a Reply