AMES, Iowa – The coronavirus that causes COVID-19 has demonstrated a stubborn ability to resist most nucleoside antiviral treatments, but a new study led by an Iowa State University scientist could help to overcome the virus’s defenses.
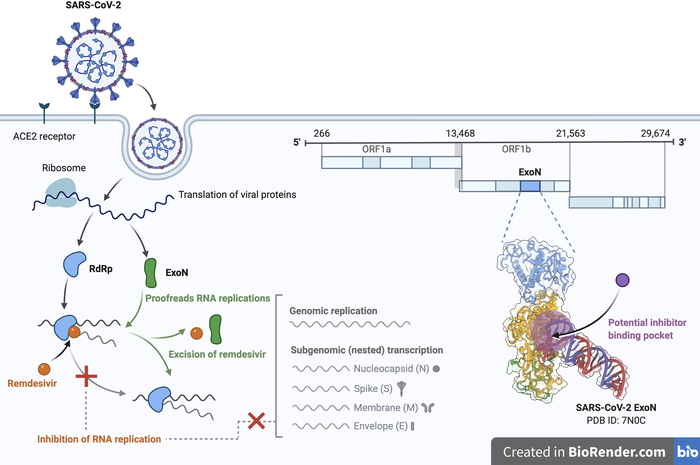
The study, published recently in the peer-reviewed journal Science, details the structure of a critical enzyme present in SARS-CoV-2, the coronavirus that causes COVID-19. This enzyme, known as the proofreading exoribonuclease (or ExoN), removes nucleoside antiviral medications from the virus’s RNA, rendering most nucleoside analogs-based antiviral treatments ineffective. The new study presents the atomic structures of the ExoN enzyme, which could lead to the development of new methods for deactivating the enzyme and opening the door to better treatments for patients suffering from COVID-19.
“If we could find a way to inhibit this enzyme, maybe we can achieve better results to kill the virus with existing nucleoside antiviral treatments. Understanding this structure and the molecular details of how ExoN works can help guide further development of antivirals,” said Yang Yang, lead author of the study and assistant professor in the Roy J. Carver Department of Biochemistry, Biophysics and Molecular Biology at Iowa State University.
SARS-CoV-2 is an RNA virus, which means its genetic material is composed of ribonucleic acid. When the virus replicates, it must synthesize RNA. But the virus’s genome is unusually large when compared to other RNA viruses, which creates a relatively high likelihood that errors arise during RNA synthesis. These errors take the form of mismatched nucleotides, and too many errors can prevent the virus from propagating.
But the ExoN enzyme acts as a proofreader, recognizing mismatches in the virus RNA and correcting errors that occur during RNA synthesis, Yang said. The enzyme is present only in coronaviruses and a few other closely related virus families, he said.
The same process that eliminates replication errors also eliminates antiviral agents delivered by the treatments commonly used to fight other RNA viruses, such as HIV, HCV and Ebola virus, which partially explains why SARS-CoV-2 has proven so difficult to treat, Yang said.
But Yang and his colleagues utilized cryogenic electron microscopy, a technique in which samples are flash cooled to cryogenic temperatures in vitreous ice to preserve their native structures, to detail the structure of the enzyme. Understanding that structure could allow for the development of molecules that bind to the enzyme and disable it. Yang said that’s the next step for his laboratory and his colleagues. Finding such a molecule could make the virus more susceptible to newly developed antivirals, Yang said. Or, it could allow for the optimization of current antivirals, such as Remdesivir.
Scott Becker, an ISU graduate student in the Roy J. Carver Department of Biochemistry, Biophysics and Molecular Biology, also contributed to the study. The study also included scientists from Yale University and the Hormel Institute at the University of Minnesota.
IMAGE CREDIT: (ENTER NAMES)



Leave a Reply