One of the big evolutionary questions in life is how and why single cell organisms organised themselves to live in a group, thereby forming multicellular life forms.
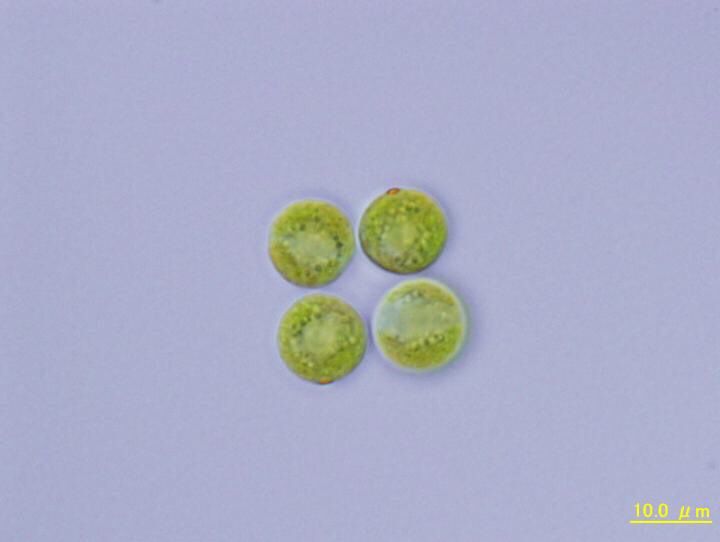
Wits PhD student, Jonathan Featherston, of the Evolution of Complexity Laboratory at the University of the Witwatersrand, Johannesburg, has answered at least part of this question, by decoding the genomic sequence of one of the simplest of all multicellular organisms – the four-celled alga Tetrabaena socialis. His research has been published in Molecular Biology and Evolution.
Tetrabaena is a member of a lineage of green-algae known as the volvocine lineage. The lineage is a model lineage for understanding how multicellularity evolved. By studying the genome of this simple alga, a number of genetic mechanisms that control how cells divide were associated with the origin of multicellularity.
By painstakingly piecing together the whole genome sequence for the alga over a period of over two years, using various genome-sequencing methodologies, Featherston has identified the ubiquitin proteasomal pathway (UPP) as a process that plays a key role in the evolution of multicellularity. This pathway is involved in regulating many activities in cells by targeting proteins for destruction thereby maintaining a careful balance of proteins in cells.
“The UPP has been implicated in many human cancers and even as a potential target for treating cancers. From this study it seems that alterations to this pathway were important for how multicellularity evolved in these algae,” says Featherston.
UPP is a complicated pathway that controls the cellular concentration of key proteins that drive cell division and it plays a role in many cellular functions. Featherston’s study suggests that UPP may play a regulating how many divisions each species of volvocine undergoes through degradation of key molecules that control cell division.
“One of the earliest evolutionary adaptations in the volvocines was a modified cell cycle. The multicellular volvocines evolved a genetic program for controlling the number of divisions during reproduction where each species has a genetically programmed maximum number of divisions. Some will only divide twice during reproduction while others may divide 12 times,” he says. “Normally people look a lot at how much of a key regulatory molecule is produced by a cell but here the interest is in the pathway that destroys these molecules. It’s kind of the other side of how cellular processes are regulated.”
Featherston compared the genome sequence of multicellular algae to their nearest single celled relative, in order to establish the genetic differences associated with the evolution of multicellularity. While overall, the the single celled and multicelled algae are very similar, he identified a small set of gene families (131) that were gained at the origin of multicellularity.
“We picked up some trends from this set of families. Many have developmental functions, which indicates that they probably are important for the evolution of multicellularity,” he says.
Featherston’s work shows that the evolution of multicellularity is associated with lineage-specific genetic developments.
“Multicellularity has evolved at least 25 times independently, but in all likelihood while certain general biological mechanisms – like cells-sticking together or modified cell cycles – may be shared, the actual genes driving these developments will mostly be unique to each lineage,” he says. “Almost all the families that are found in other organisms can be found in a diverse array of unicellular organisms, suggesting that the genes that gave rise to multicellularity were derived from genes that were already present in the unicellular ancestor but may have been duplicated to form new genes that now have new functions.”
IMAGE CREDIT: Hisayoshi Nozaki and Yoko Arakaki




Leave a Reply