HAVE YOUR SAY.
We are proud to announce the inaugural session of The Bullpen, where the members of the Scientific Inquirer community get to shape the site’s editorial decision making. We’ll be discussing people and companies to profile on the site. On Wednesday June 8th at 5:30pm EST, join us on Discord and let’s build the best Scientific Inquirer possible.
Researchers in Japan have discovered that the local DNA motion inside of human cells remains steady throughout interphase, where the cell grows and replicates its DNA for cell division. The study suggests that this steady-state DNA motion allows cells to conduct housekeeping tasks under similar environments during interphase.
The team, led by Professor Kazuhiro Maeshima of National Institute of Genetics, ROIS, published their findings June 3 in Science Advances.
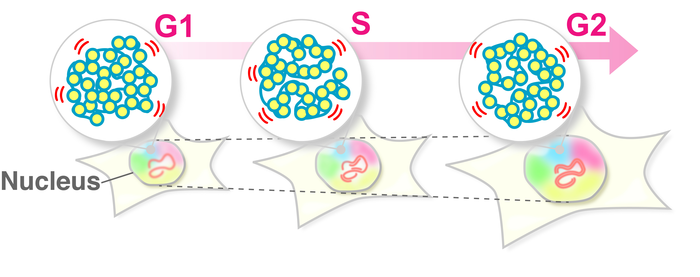
To fit inside the nucleus of the cell, DNA is organized into chromatin, in which the strands of DNA are wrapped around groups of histone proteins, like thread around a spool, to form structures known as nucleosomes. Nucleosomes can then be folded up into even more compact structures and form chromatin. Previous research shows that chromatin is continuously swaying in living cells.
As the cell cycle progresses (namely G1, S, and G2 phases), where genome DNA doubles and the nucleus becomes larger, the nuclear environment surrounding chromatin drastically changes. Maeshima and colleagues at the National Institute of Genetics in Mishima, Japan, constructed this question: How does chromatin behavior change during interphase?
Maeshima’s group used a high-resolution light microscopy technique to look at the behavior of individual nucleosomes inside living cells for a very short time, approximately one second.
Maeshima and colleagues have revealed that the local chromatin motion remains steady throughout interphase, although genome DNA is doubled by DNA replication and the nucleus grows. The researchers also have shown that nuclear growth without replication did not affect the steady-state motion of chromatin. Thus, local chromatin motion is independent of such nuclear changes during interphase.
“This is an important finding because the steady-state motion allows cells to conduct their routines, such as RNA transcription and DNA replication, under similar nuclear environments,” the first author Shiori Iida said. “Local chromatin motion can govern genomic DNA accessibility for target searching or recruiting a piece of machinery. The steady-state motion of chromatin provides a robust cellular system in which DNA functions are unaffected by various nuclear changes.”
“Cells can transiently change the chromatin motion from the steady state to perform their ad hoc jobs in response to DNA damages, among many other tasks,” Maeshima said. He and his team aim to further explore how DNA motion is regulated, which proteins are involved in the regulation process, and more about how DNA behaves during cell division. “Our ultimate goal is to understand how human genomic DNA inside the cell behaves to read-out genetic information in it,” Maeshima said.
IMAGE CREDIT: Shiori Iida & Kazuhiro Maeshima, National Institute of Genetics, ROIS


Leave a Reply