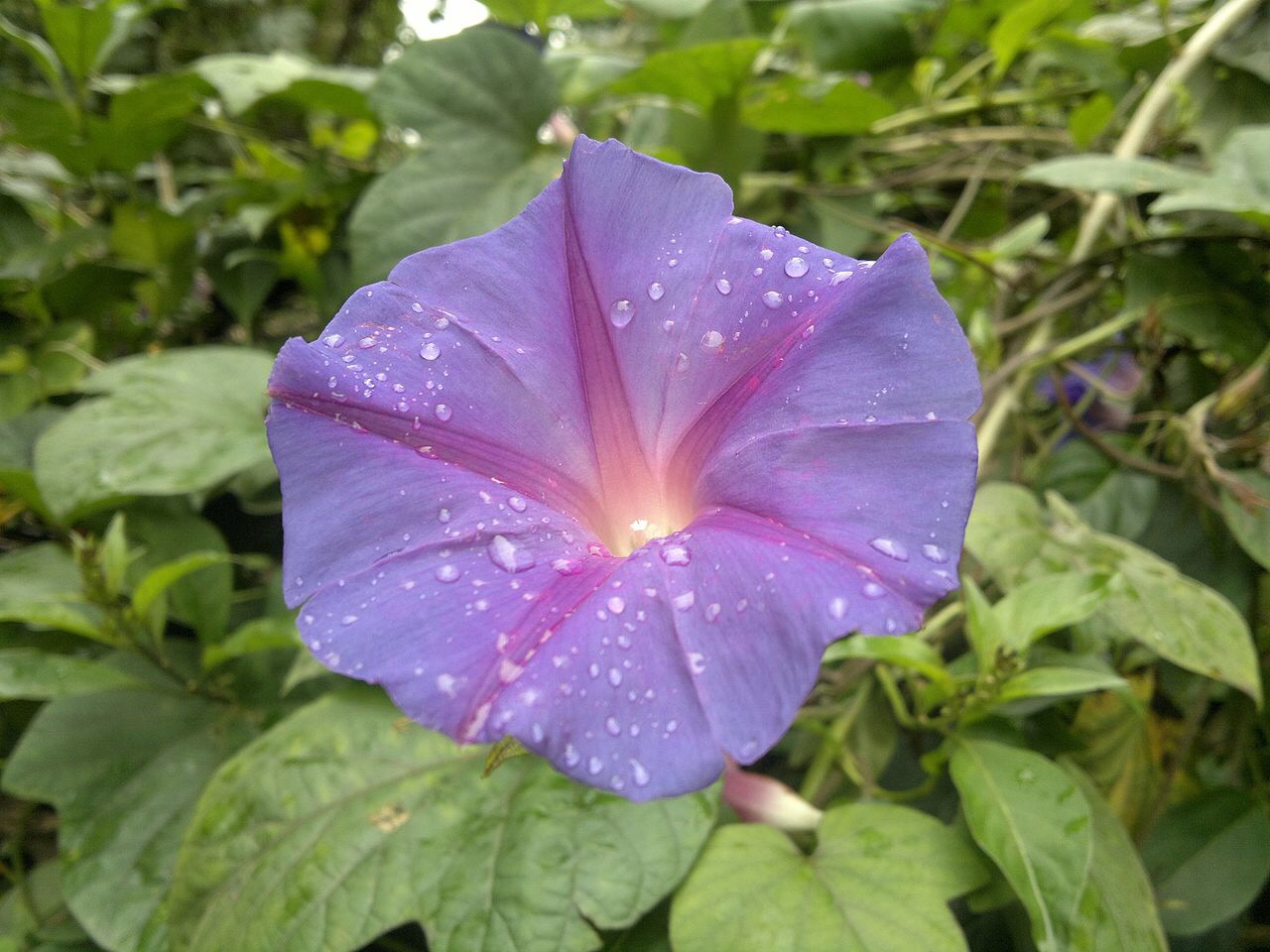
In a world-first, Japanese scientists have used the revolutionary CRISPR, or CRISPR/Cas9, genome- editing tool to change flower colour in an ornamental plant. Researchers from the University of Tsukuba, the National Agriculture and Food Research Organization (NARO) and Yokohama City University, Japan, altered the flower colour of the traditional Japanese garden plant, Japanese morning glory (Ipomoea nil or Pharbitis nil), from violet to white, by disrupting a single gene. This research highlights the huge potential of the CRISPR/Cas9 system to the study and manipulation of genes in horticultural plants.
Japanese morning glory, or Asagao, was chosen for this study as it is one of two traditional horticultural model plants in the National BioResource Project in Japan (NBRP). Extensive genetic studies of this plant have already been performed, its genome sequenced and DNA transfer methods established. In addition, as public concern with genetic technologies such as CRISPR/Cas9 is currently a social issue in Japan, studies using this popular and widely-grown plant may help to educate the public on this topic.
The research team targeted a single gene, dihydroflavonol-4-reductase-B (DFR-B), encoding an anthocyanin biosynthesis enzyme, that is responsible for the colour of the plant’s stems, leaves and flowers. Two other, very closely related genes (DFR-A and DRF-C) sit side-by-side, next to DFR-B. Therefore, the challenge was to specifically and accurately target the DFR-B gene without altering the other genes. The CRISPR/Cas9 system was used as it is currently the most precise method of gene editing.
The CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats)/Cas9 system is based on a bacterial defense mechanism. It is composed of two molecules that alter the DNA sequence. Cas9, an enzyme, cuts the two strands of DNA in a precise location so that DNA can be added or removed. Cas9 is guided to the correct location by gRNA, or guide RNA, a small piece of RNA that has been designed to be complementary to the target DNA sequence. Cas9 cuts the two strands of DNA at the target location, allowing DNA to be removed and/or added.
As reported on 30 August 2017 in Scientific Reports, a short DNA sequence in the Japanese morning glory DFR-B gene was selected as the target for the CRISPR/Cas9 system. This sequence contains the active site of the enzyme produced by the DFR-B gene. Disruption of this sequence should therefore de-activate the enzyme, resulting in an absence of the colour pigment, anthocyanin.
The CRISPR/Cas9 system was inserted into tissue-cultured embryos of Japanese morning glory plants using the DNA-transferring capabilities of the plant bacterium Rhizobium. As expected, the DFR-B enzyme was successfully inactivated, resulting in approximately 75%of the transgenic plants with green stems and white flowers. Non-transformed plants with an active enzyme had violet stems and flowers. These changes in stem colour were observed very early in the tissue culture process.
A series of genetic analyses confirmed that the DNA target sequence had been altered in the transgenic plants, with either DNA insertions or deletions in both copies of the DFR-B gene (so-called bi-allellic mutants). The other related genes, DFR-A and DFR-C, were examined and no mutations were found, confirming the high specificity of the CRISPR/Cas9 system.
Next, the researchers examined the inheritance of the CRISPR/Cas9-induced mutations by analyzing plants from the next generation. These plants looked exactly like their parents. Among these plants were some without any sign of the introduced DNA. This raises some interesting questions in terms of the regulation of genetically modified organisms (GMOs), as these next-generation plants are considered transgenic, based on process-based definitions (how they were made), and non- transgenic, based on product-based definitions (the presence of foreign DNA in the final product).
This technology is also extremely useful in confirming the function of genes. Experiments in the 1980s and 1990s used ‘forward’ genetic screening techniques to find the genes responsible for flower colour production in the Japanese morning glory. The CRISPR/Cas9 system described here is the ‘reverse’ genetic approach, used to find out what an organism looks like after a known gene is disrupted, and confirms that the DFR-B gene is the main gene responsible for colour in Japanese morning glory plants.
Currently, CRISPR/Cas9 technology is not 100% efficient, that is, not all targeted plants will be transgenic. The mutation rate in this study, 75%, however, was relatively high. This is one of the reasons this research will greatly facilitate those interested in the modification of flower colours and shapes using the CRISPR/Cas9 system in ornamental flowers or vegetables.
The story of the Japanese morning glory started in the 8th century AD, with the introduction of wild blue-flowered plants into Japan from China. In 1631, the first white-flowered Japanese morning glory was painted in Japan. What took nature nearly 850 years to achieve has taken less than one using the CRISPR/Cas9 system, indicating both its power and its potential.



Leave a Reply